Difference between revisions of "Summary"
(→Screenshots) |
(→Screenshots) |
||
| Line 62: | Line 62: | ||
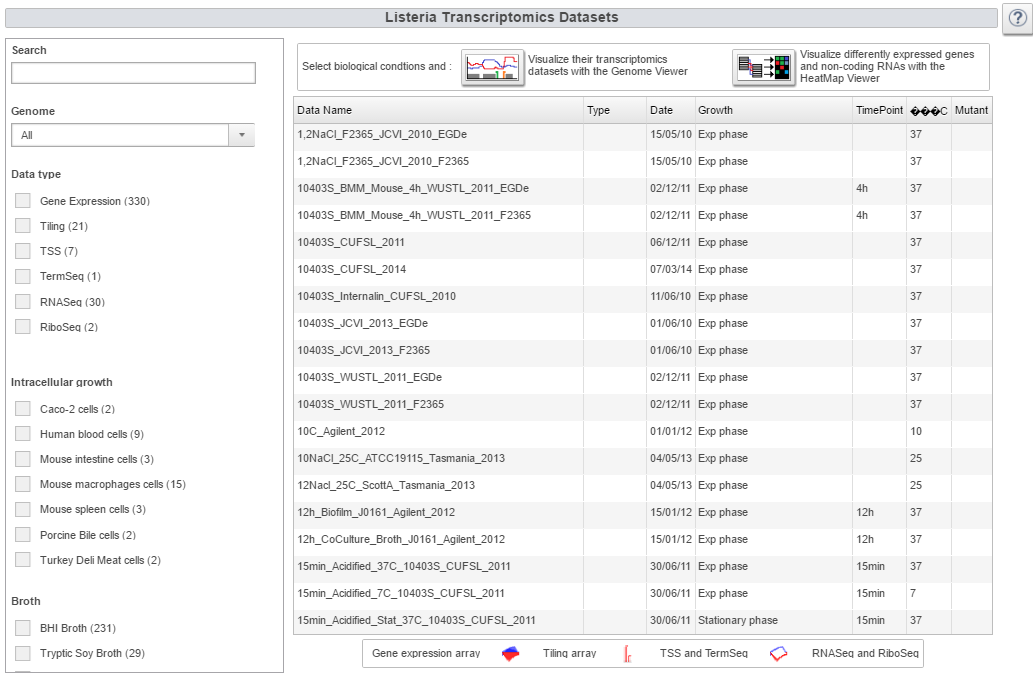
Image:transcriptome.png|Transcriptome webpage | Image:transcriptome.png|Transcriptome webpage | ||
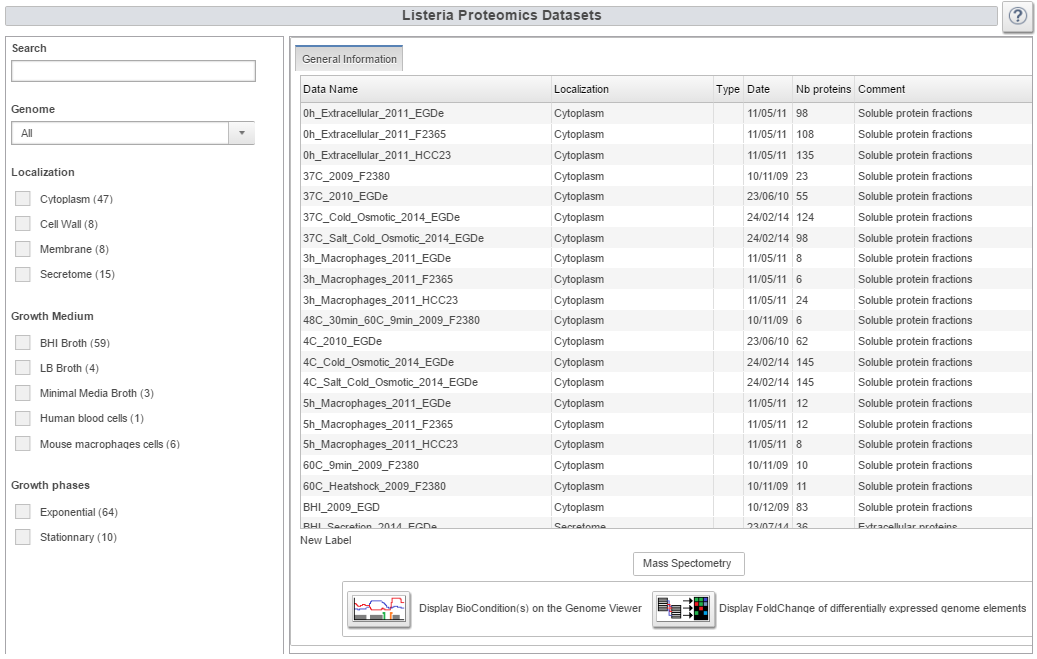
Image:proteome.png|Proteome webpage | Image:proteome.png|Proteome webpage | ||
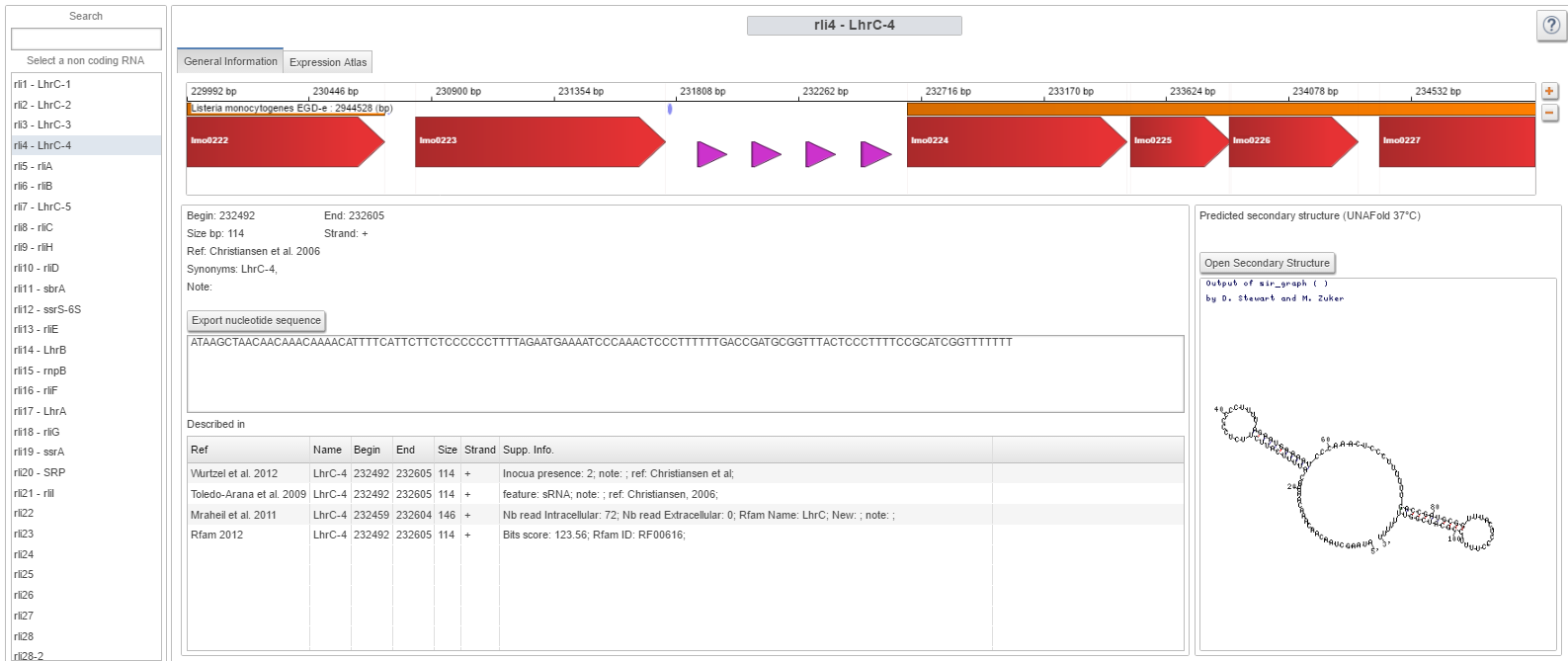
| + | Image:SeqInfo.png|Sequence information panel | ||
Image:coexpression.png|Co-expression network webpage | Image:coexpression.png|Co-expression network webpage | ||
</gallery> | </gallery> | ||
Revision as of 10:43, 19 October 2016
Listeriomics integrates all complete genomes, transcriptomes and proteomes published for Listeria species to date. It allows navigating among all these datasets with enriched metadata in a user-friendly format. Use Listeriomics for deciphering regulatory mechanisms of your genome element of interest
The main purpose of the website is to give scientists a quick and easy access to tools created for answering the four questions one has when starting a study on a specific genomic element:
- What are the known functions of a genomic feature in a given strain and homologies in closely related strains?
- What are the biological conditions in which a genomic feature is transcribed?
- What are the biological conditions in which a genomic feature is translated?
- What is the regulation network involved with a genomic feature of interest?
- What do I know about my gene/small RNA function, and is it present in other strains?
- Is my genomic element transcribed, and in which biological condition?
- Is my genome element translated, and in which biological condition can I find the protein?
- Is my gene/small RNA a regulator or does it regulate another genomic element?
The website integrates different type of tools for “omics” data analyses:
- A highly interactive genome viewer for display of gene expression arrays, tiling arrays, and RNA-Seq datasets along with proteomics and genomics datasets.
- Expression and protein atlas, which are tools to connect every genomic feature (genes, smallRNAs, antisenseRNAs) or proteins to the most relevant “omics” data.
- A tool to explore gene conservation throughout Listeria strains.
- A co-expression network tool to discover new potential regulation.
Tutorials
| First hands on Listeriomics | General organization of the website and principal features |
| Genomic tools | How-to access all the genomes. |
| Gene tools | How-to access a specific gene and get information about its annotation, its conservation, its transcription or translation. |
| Small RNA tools | How-to access a specific Small RNA |
| Transcriptomic tools | How-to access transcriptomic dataset and display it or perform an analysis with the expression atlas |
| Proteomic tools | How-to access proteomic dataset and display it or perform an analysis with the protein atlas |
Database
The Listeriomics database includes all complete genomes, transcriptomes and proteomes published on Listeria species to date. It allows scientists to analyse dynamically all these datasets in a user-friendly format, and can be used by bio-informaticians as a central database for systems biology analysis.
List of data available:
- 83 Listeria complete genomes
- 492 transcriptome datasets
- 74 proteome datasets