Small RNA tools
Contents
General organization[edit]
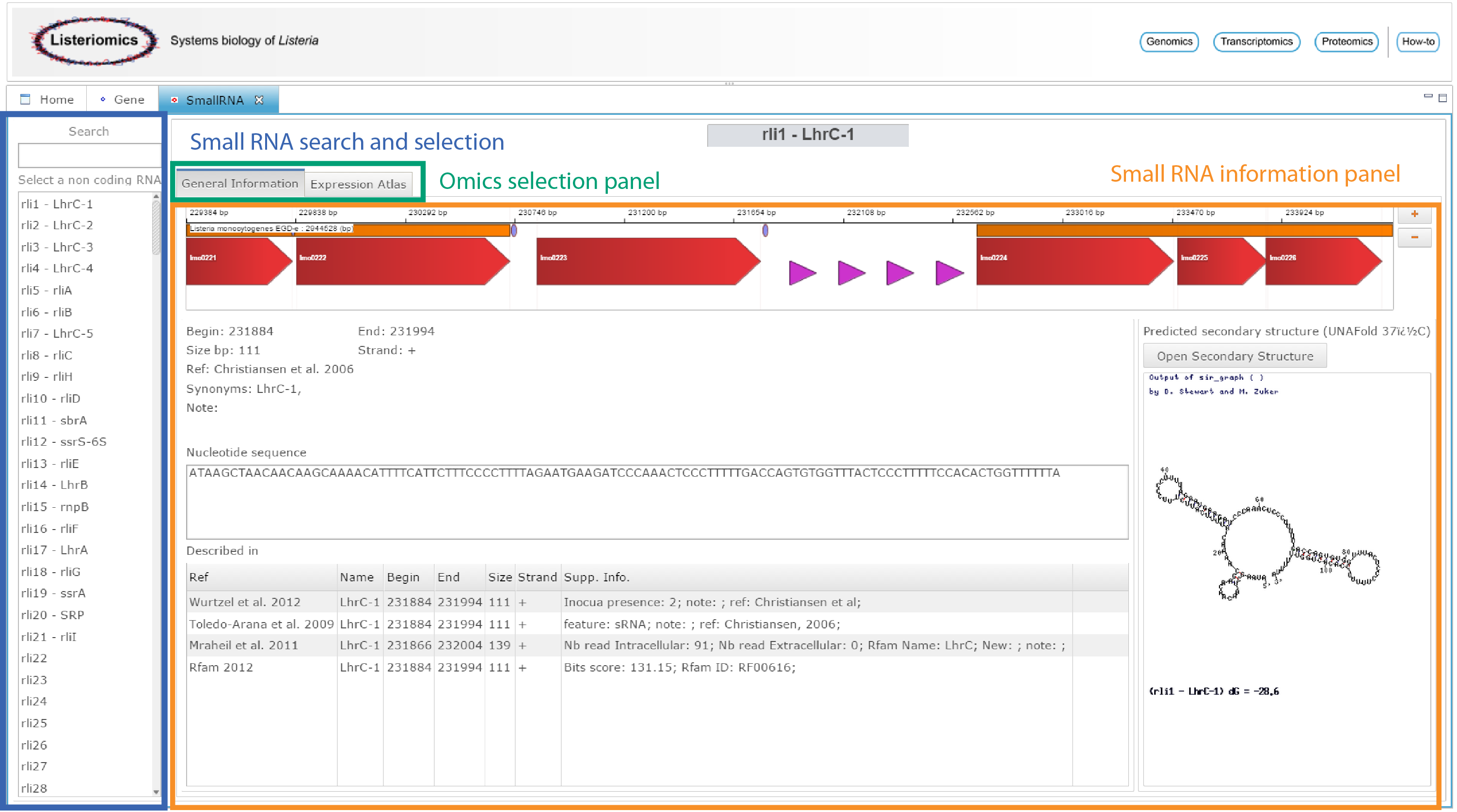
The small RNA webpage can be accessed either from the home webpage of the Listeriomics website or by double-clicking on a small RNA in the genome viewer.
| Small RNA search and selection
In this list, all the small RNA available in Listeria monocytogenes EGD-e strain are displayed. One can search for a small RNA. A click on an element of the list will display all related information on the Small RNA information panel. |
| Omics selection panel
For each Listeria small RNA two different information panels are available.
|
| Small RNA information panel
The panel in which all information will be displayed |
How-to access Small RNA information ?[edit]
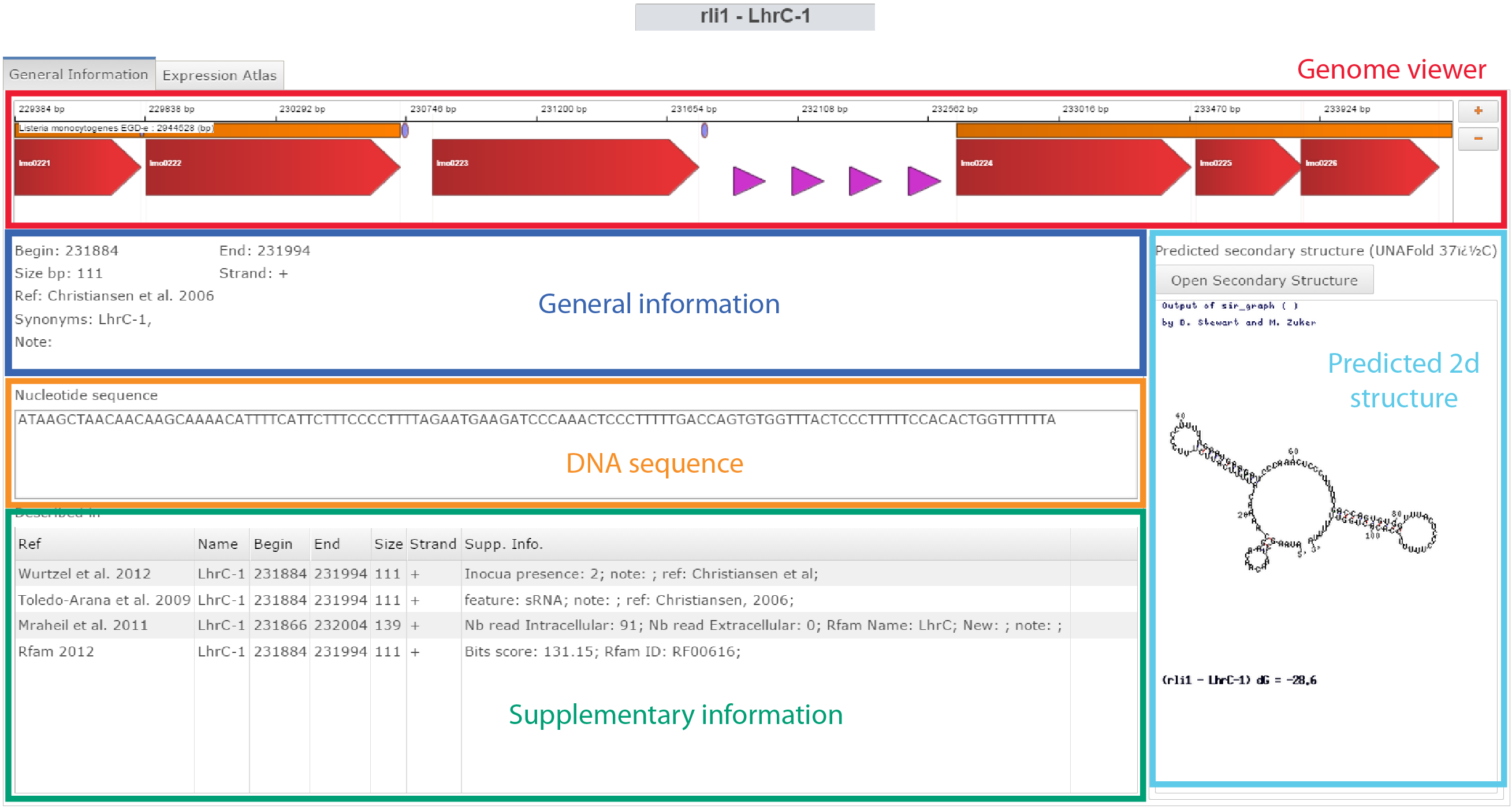
Click on General information on the Omics selection panel
| |
| DNA sequences can be accessed here and saved to fasta file or sent directly to Blast. |
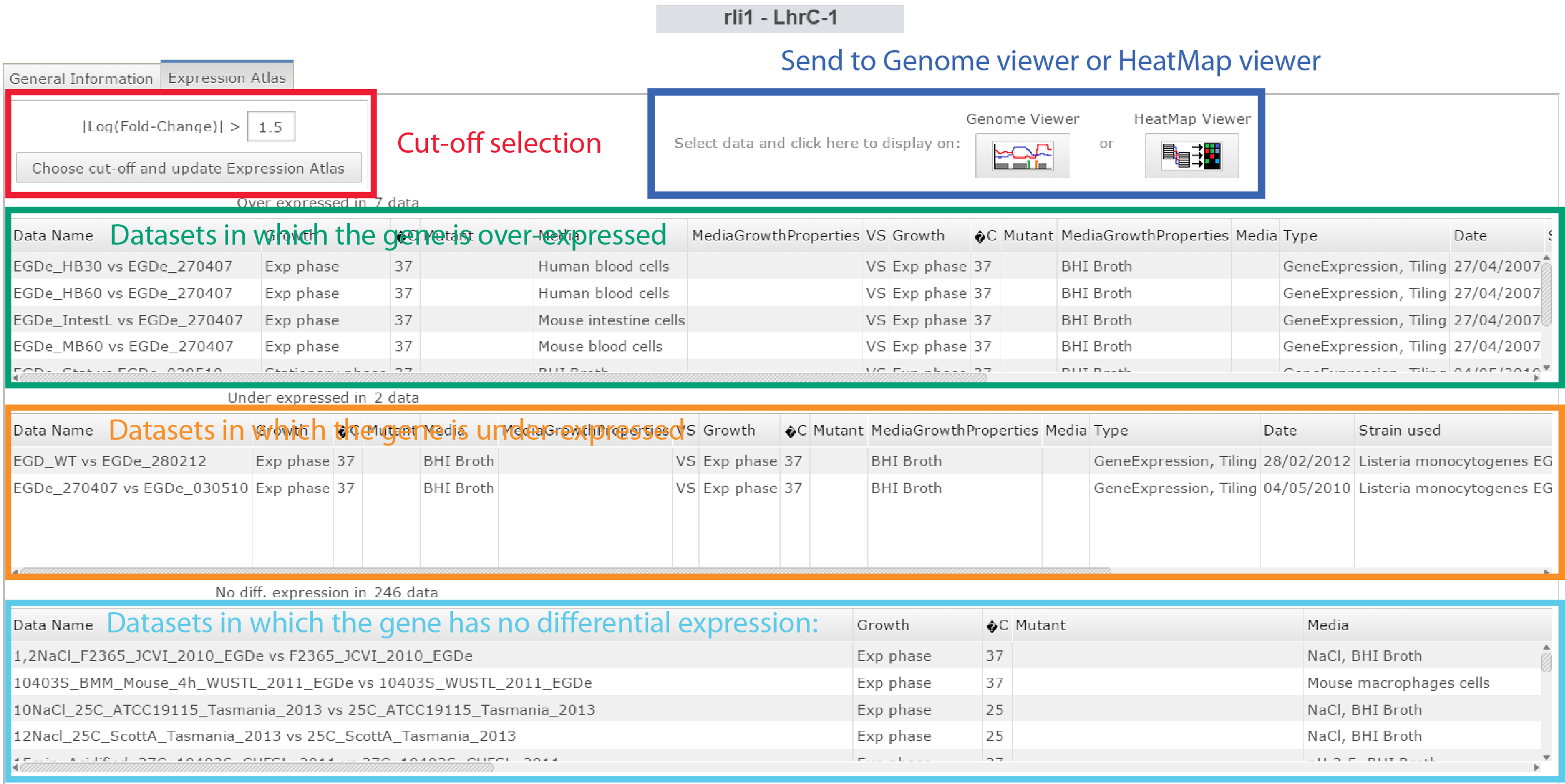
Is my Small RNA transcribed ?[edit]
Click on Expression Atlas on the Omics selection panel
|
Summary of Small RNA studies in Listeria monocytogenes EGD-e[edit]
In the past decade, noncoding RNAs in Listeria have been studied in detail [1] [2] [3] [4] [5] [6].
One study concerns strain 10403S [4], and all the other studies concern the EGD-e strain.
Altogether, 305 noncoding RNA elements are reported in Listeria monocytogenes EGD-e. They are separated in three groups:
- 155 sRNAs (acting mainly in trans)
- 46 cis regulatory elements (cisRegs)
- 104 antisense RNAs (asRNAs)
Tables summarizing all the small RNAs observed can be accessed in the small RNAs summary webpage. This page can be accessed from the home webpage of the Listeriomics website.
| Small RNA type selection:
Type of small RNA to display:
|
| Small RNAs summary table
Table with different information on each small RNA:
By clicking on a small RNA, the small RNA webpage will open to give more information. |
References[edit]
Template:References- ↑ Christiansen JK, Nielsen JS, Ebersbach T, Valentin-hansen P, Søgaard- Andersen L, Kallipolitis BH. 2006. Identification of small Hfq-binding RNAs in Listeria monocytogenes. RNA 12:1383–1396.
- ↑ Mandin P, Repoila F, Vergassola M, Geissmann T, Cossart P. 2007. Identification of new noncoding RNAs in Listeria monocytogenes and pre- diction of mRNA targets. Nucleic Acids Res. 35:962–974.
- ↑ Toledo-Arana A, Dussurget O, Nikitas G, Sesto N, Guet-Revillet H, Balestrino D, Loh E, Gripenland J, Tiensuu T, Vaitkevicius K, Bar- thelemy M, Vergassola M, Nahori MA, Soubigou G, Régnault B, Cop- pée JY, Lecuit M, Johansson J, Cossart P. 2009. The Listeria transcrip- tional landscape from saprophytism to virulence. Nature 459:950–956.
- ↑ 4.0 4.1 Oliver HF, Orsi RH, Ponnala L, Keich U, Wang W, Sun Q, Cartinhour SW, Filiatrault MJ, Wiedmann M, Boor KJ. 2009. Deep RNA sequencing of L. monocytogenes reveals overlapping and extensive stationary phase and sigma B-dependent transcriptomes, including multiple highly transcribed noncoding RNAs. BMC Genomics 10:641.
- ↑ Mraheil MA, Billion A, Mohamed W, Mukherjee K, Kuenne C, Pischi- marov J, Krawitz C, Retey J, Hartsch T, Chakraborty T, Hain T. 2011. The intracellular sRNA transcriptome of Listeria monocytogenes during growth in macrophages. Nucleic Acids Res. 39:1–14.
- ↑ 6.0 6.1 Wurtzel O, Sesto N, Mellin JR, Karunker I, Edelheit S, Bécavin C, Archambaud C, Cossart P, Sorek R. 2012. Comparative transcriptomics of pathogenic and non-pathogenic Listeria species. Mol. Syst. Biol. 8:1–14. [http://dx.doi.org/10.1038/msb.2012.11.