Proteomic tools
General organization[edit]
The proteomic tool can be accessed by clicking on the button on the right of the banner of the Listeriomics website.
We extracted 102 proteomics files (74 absolute value datasets and 28 relative expression datasets)
| Filter panel
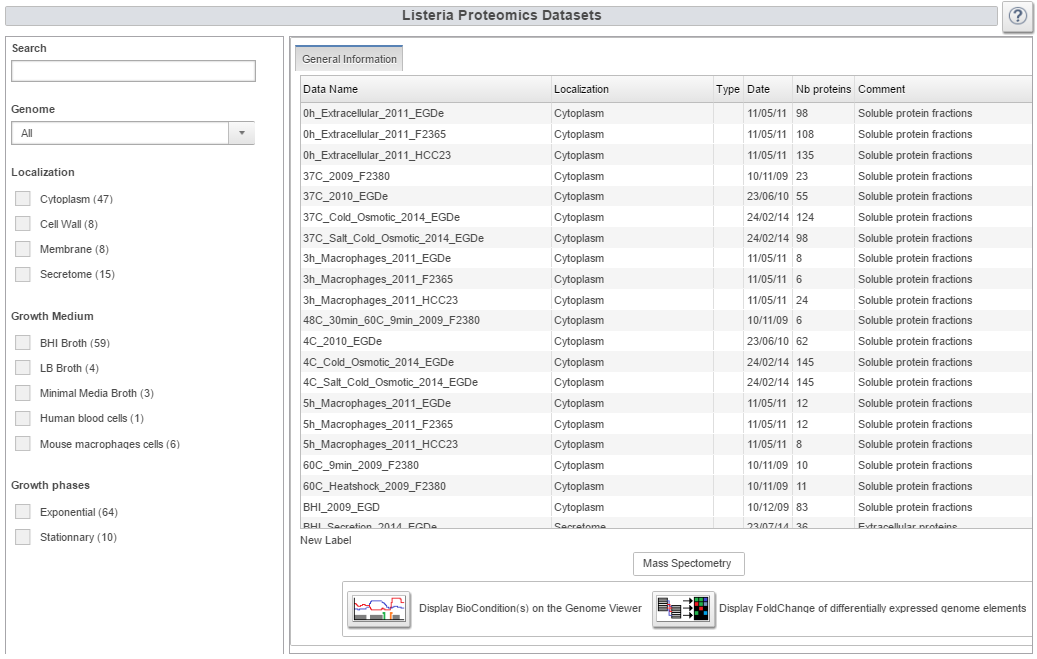
With this panel one can filter out the list of proteomics datasets by different key properties: Genome, Data type, ... A search bar is available to do so, or by clicking on the check box. In parenthesis is displayed the number of datasets available for each property. |
| List of all Listeria proteomic datasets with biological condition information
All proteomic datasets available are displayed with all additional information: “TimePoint” for growth time; “Mutant” if a gene is removed or over-expressed; “Growth” indicates the growth phase from which the bacteria were extracted; “Temperature” for the growth media; “Media” gives the media properties for example if Glucose was added, or if bacteria were extracted from macrophages; “Strain used” indicates the strain used for the experiment. [IGNOU Hall ticket][1] |
| Link to Genome and Heatmap viewers
If proteomic datasets are selected in the summary table, there are two possibilities: Display them on the genome viewer for direct visualization, or look at each dataset in the heatmap viewer. |
Proteomic database[edit]
Since no centralized database like ArrayExpress exists to browse proteomics data, we selected a list of 23 publications using the PubMed database in which a mass spectrometry experiment with a Listeria strain was performed. From core articles and supplementary files we extracted all available information on the biological conditions screened in each experiment. We extracted the information in the same format as for the transcriptomic datasets. In all the experiments a list of proteins detected was available, and in some cases we retrieved a list of proteins differently produced between two biological conditions.